Summary
A research team at Vanderbilt University Mass Spectrometry Research Center has developed the Molecular Image Fusion software system, that by fusing spatial correspondence between histology and imaging mass spectrometry (IMS) measurements and cross-modality modeling, can predict ion distributions in tissue at spatial resolutions that exceed their acquisition resolution. The prediction resolution can even exceed the highest spatial resolution at which IMS can be physically measured. This software has been successfully tested on different IMS datasets and can be extended to other imaging modalities like MRI, PET, CT, profilometry, ion mobility spectroscopy, and different forms of microscopy.
Addressed Need
A typical IMS experiment collects both IMS measurements and stained microscopic images of the tissue section or a matching serial tissue section. Current state-of-the-art in examining cross-modality patterns in molecular imaging modalities is to overlay images coming from different imaging sources and to leave the recognition of any cross-modality patterns and relationships to the human observer, which may be time consuming, costly, and prone to errors. There is no suitable mapping software to accurately map imaging modalities across the platforms and measurement technologies.
Technology Description
The Molecular Image Fusion software tool consists of a system that takes the measurements made by two or more molecular imaging modalities and exhaustively captures the cross-correlating patterns and relationships between these observations in a rigorous mathematical model. Once this cross-modality model has been established, the system uses the model to achieve one or more predictive, preprocessing, filtering, or discovery tasks across multiple data sources.
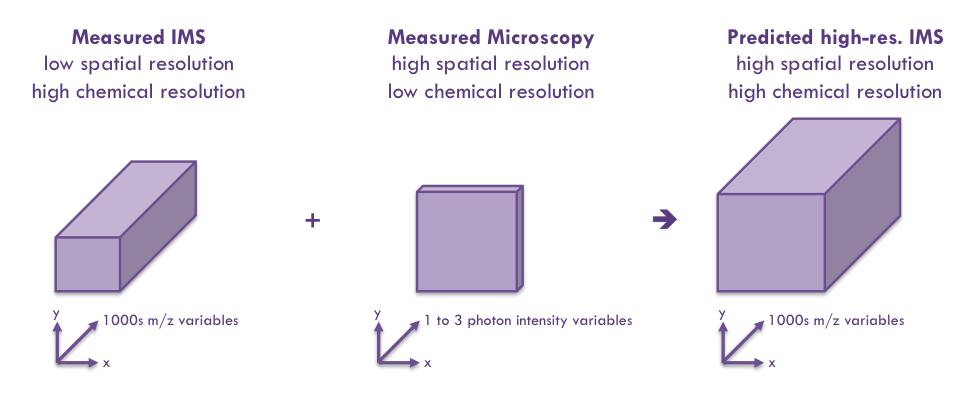

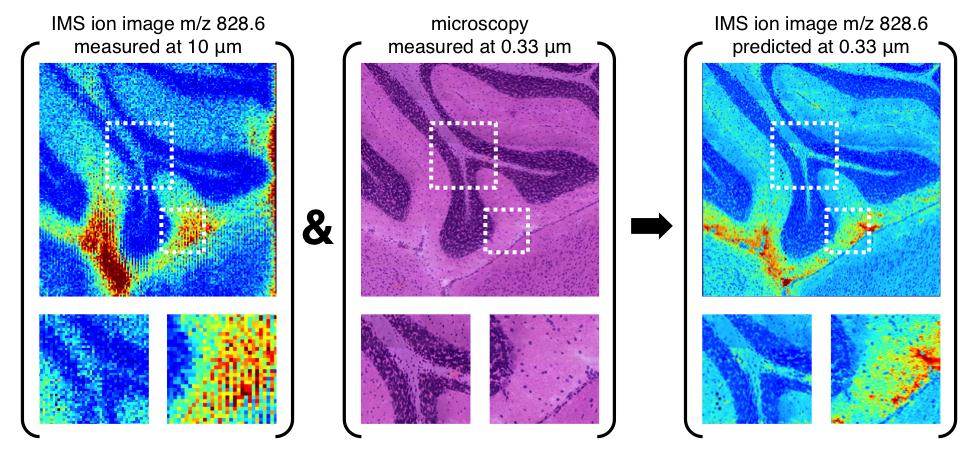
Molecular Image Fusion software predicts ion distributions in tissue at spatial resolutions that exceed their acquisition resolution. Picture shows a sub-micron prediction of the distribution of ion m/z 828.6 in mouse brain, achieved by Molecular Image Fusion using H&E stained microscopy at 0.33 μm and an IMS measurement on a neighboring tissue section acquired at 10 μm.
Applications
Predictions of IMS values are applicable where conducting a physical high-resolution IMS measurement is unpractical (e.g. measurement time, instrument stability), uneconomical (e.g. laser wear, detector deterioration), or even infeasible (e.g. spatial resolution below laser bandwidth, low SNR).
Additionally, image fusion allows for modality-specific patterns to be separated from patterns corroborated across the modalities, benefitting both low-level preprocessing (e.g. de-noising) and high-level interpretation (e.g. increasing confidence in biological and sample patterns supported by multiple measurement principles).
Technology Development Status
The source code for the proof-of-concept method implementation using various test examples has been developed. Ongoing development of the software for applications in MRI, PET, CT, profilometry, ion mobility spectroscopy modalities, and for different forms of microscopy (e.g. phase, DIC).
Intellectual Property Status
Provisional patent application filed in 2012.
Research group webpage and list of publications: Vanderbilt University Mass Spectrometry Research Center; http://www.vanderbilt.edu/chemistry/faculty/caprioli.php
VU Reference: VU12169




